TLDR
TDP-43 is a protein that normally helps regulate genes in the nucleus, but in ALS and frontotemporal dementia it mislocalizes to the cytoplasm and forms toxic clumps. The M337V mutation, classified as pathogenic and not found in healthy populations, causes this protein to misfold with moderate structural confidence (64.8% average). This mutation triggers early cellular dysfunction including impaired energy production and axonal transport even before visible protein aggregates form, helping explain how it causes motor neuron death in ALS patients.
Detailed Analysis
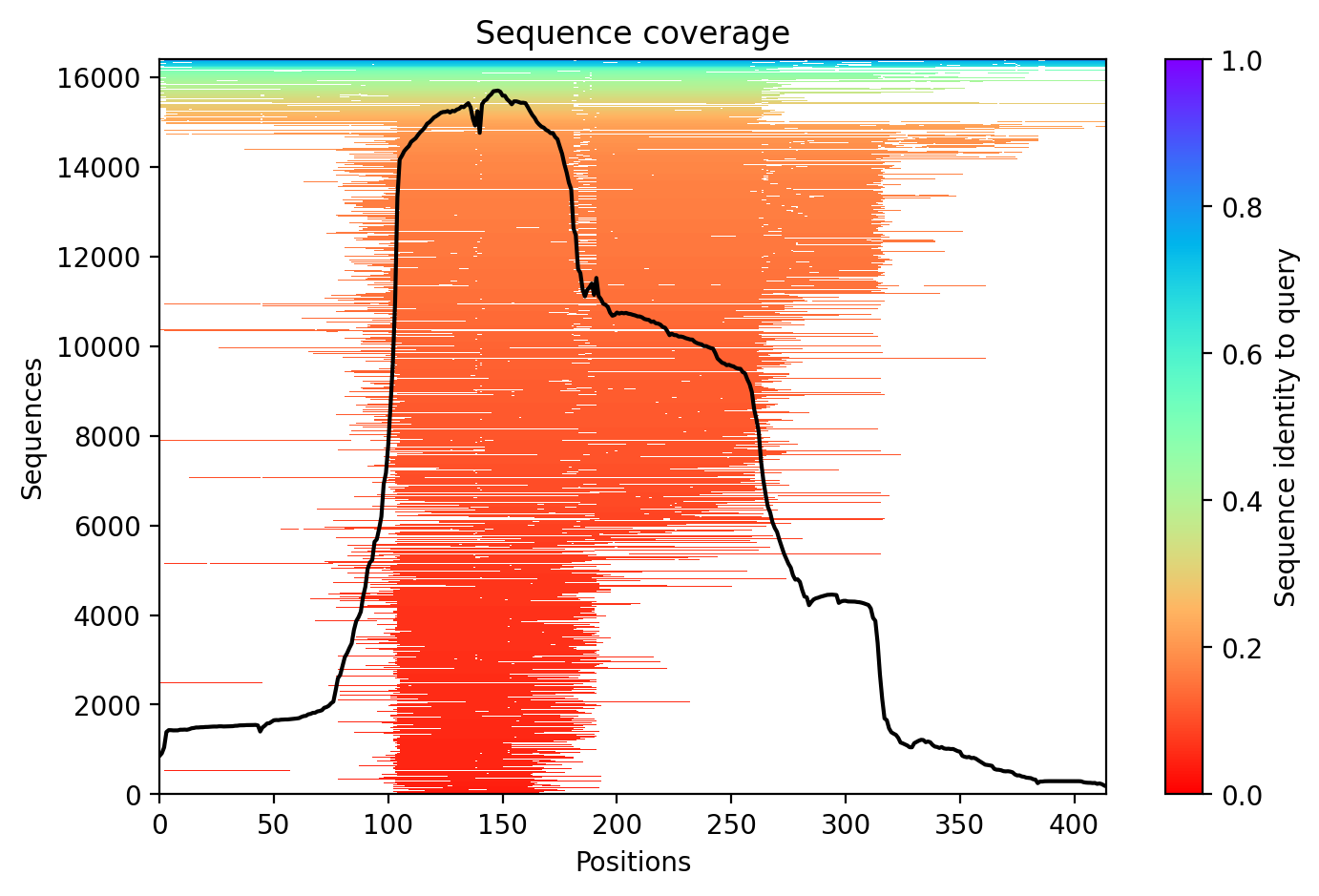
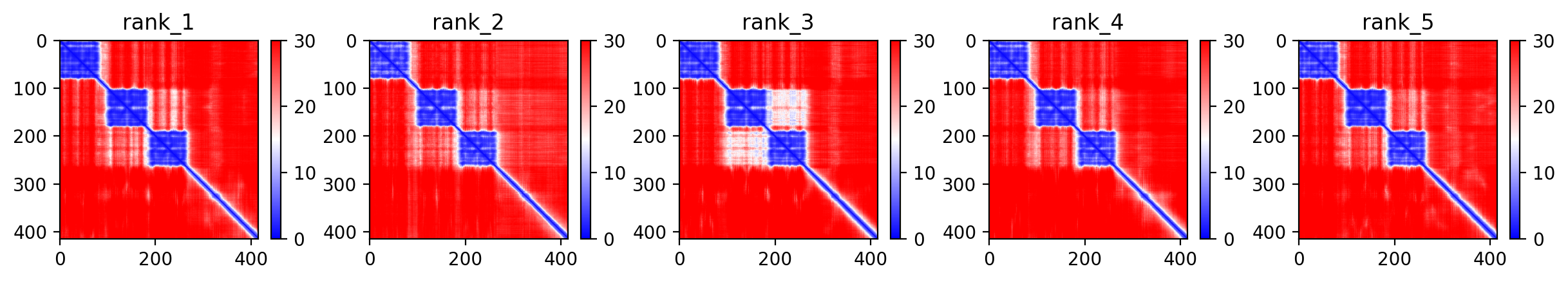
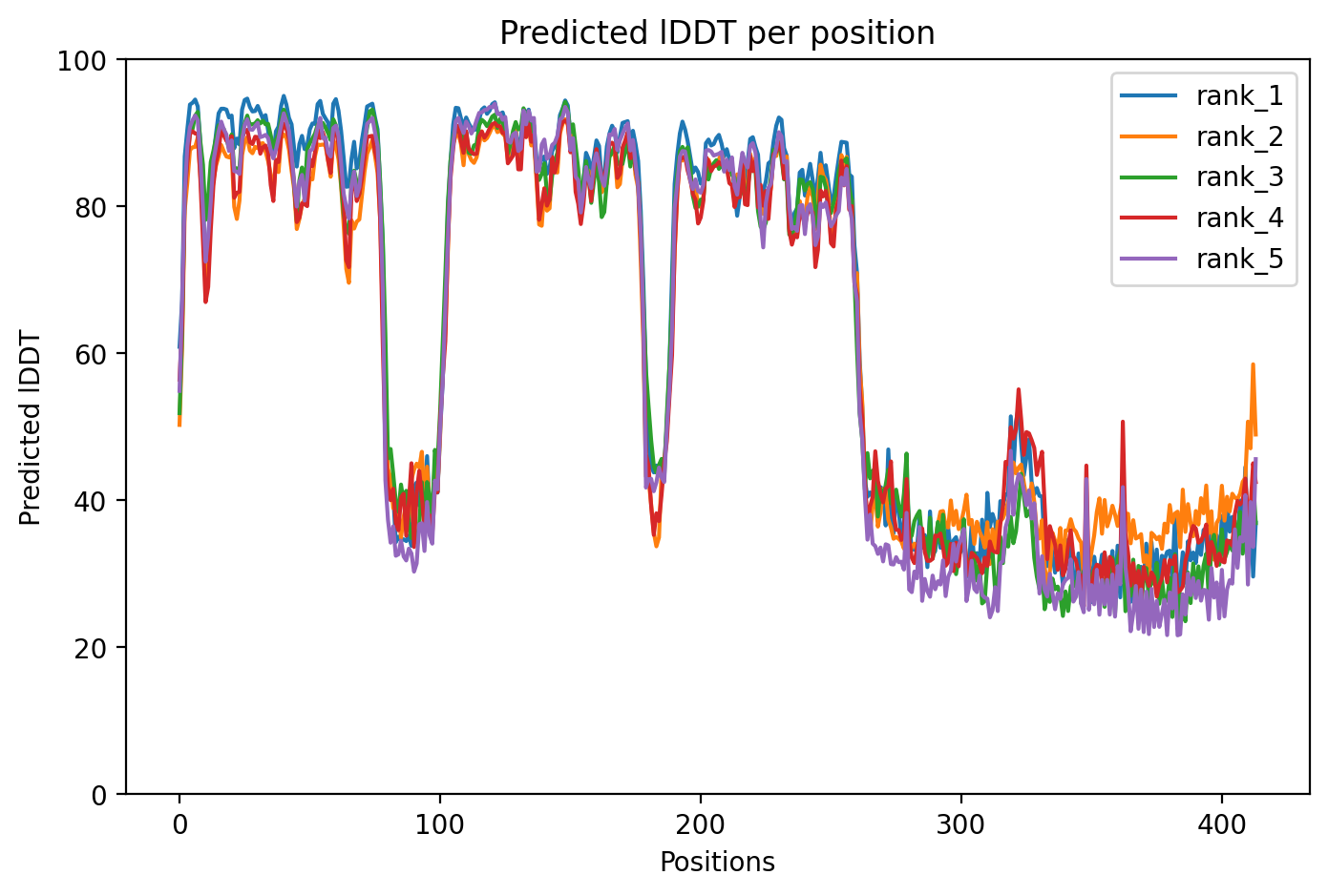
The M337V mutation in TDP-43 represents a pathogenic variant linked to both amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD), with expert classification as pathogenic/likely pathogenic and absence from healthy population databases. The AlphaFold2 structural prediction achieves a moderate average confidence score of 64.8% (pLDDT), indicating some structural uncertainty particularly in disordered regions typical of TDP-43's C-terminal domain where M337V resides. This confidence level allows for general structural insights but requires caution when interpreting fine structural details.
The M337V mutation disrupts multiple cellular functions even at low expression levels and without requiring visible protein aggregates. Studies in mouse motor neurons show that M337V impairs cell viability, reduces neurite outgrowth (the branching extensions neurons use to communicate), slows axonal transport (the cellular highway system moving materials along nerve fibers), and disrupts glycolysis (the process cells use to generate energy) under normal unstressed conditions [1][4]. In patient-derived neurons, M337V increases both soluble and detergent-resistant forms of TDP-43, elevates motor neuron death risk by nearly threefold, and makes neurons more vulnerable to loss of growth factor signaling, indicating problems with cellular survival pathways [2]. Additionally, inhibiting glycogen synthase kinase-3 (GSK3), an enzyme that M337V-containing TDP-43 inappropriately activates, can suppress neurotoxicity through caspase-dependent mechanisms (cellular suicide pathways), suggesting potential therapeutic targets [2].
The structural location of M337V in the C-terminal region has significant functional consequences. This mutation increases aggregation propensity in full-length TDP-43 protein, alters how the protein behaves in stress granules (cellular structures that form under stress), and disrupts liquid-liquid phase separation (LLPS), a process where proteins reversibly concentrate into droplets that can become pathological if they harden. Mouse models expressing M337V develop reactive gliosis (inflammatory activation of support cells), abnormal protein ubiquitination (cellular tagging for degradation), gait problems, early death, mitochondrial aggregates (clumped energy-producing structures), and phosphorylated tau protein (another hallmark of neurodegeneration). The mutation also reduces levels of the cell's own TDP-43 and related RNA-binding protein hnRNP K [1][4].
Beyond motor neurons, M337V affects multiple cell types relevant to ALS/FTD pathology. Recent work shows that endothelial cells (which form blood vessel walls) from ALS-FTD patients have reduced nuclear TDP-43, contributing to blood-brain barrier defects that may accelerate disease progression [5]. In serotonergic neurons in C. elegans worms, cytoplasmic TDP-43 causes early behavioral impairments before any neurodegeneration occurs, demonstrating that functional deficits precede cell death [1][4]. Additionally, pathology extends beyond the nervous system, with shared mechanisms including DNA damage, decreased SIRT1 levels (a protective protein), and increased acetylated p53 (a stress response protein) found across sporadic ALS and familial FTD cases, suggesting common pathways despite different genetic causes [3].
The clinical significance of M337V extends across the ALS-FTD spectrum. The mutation has been identified in patients presenting with classical ALS, pure frontotemporal dementia, and atypical presentations including facial-onset sensory and motor neuronopathy (FOSMN), where sensory loss in the face precedes motor weakness spreading down the body [6]. The pathogenic classification reflects consistent evidence that this variant disrupts protein function through multiple mechanisms: enhanced aggregation, impaired nuclear-cytoplasmic trafficking, disrupted RNA metabolism, mitochondrial dysfunction, and compromised cellular energy production. The absence of M337V in population databases strengthens the case that this is a disease-causing mutation rather than a benign variant, as truly pathogenic mutations are typically rare or absent in healthy individuals.
Works Cited
[1] Lacour et al. (2026). Cytoplasmic TDP-43 leads to early behavioral impairments without neurodegeneration in a serotonergic neuron-specific C. elegans model. Scientific reports. [PubMed](https://pubmed.ncbi.nlm.nih.gov/41571758/)
[2] White et al. (2026). Inhibiting Glycogen Synthase Kinase 3 Suppresses TDP-43-Mediated Neurotoxicity in a Caspase-Dependent Manner. Molecular neurobiology. [PubMed](https://pubmed.ncbi.nlm.nih.gov/41546756/)
[3] Jun et al. (2025). The Ku80-p53-SIRT1 axis in DNA damage response contributes to sporadic and familial ALS and FTD. Nature communications. [PubMed](https://pubmed.ncbi.nlm.nih.gov/41422089/)
[4] Lacour et al. (2025). Cytoplasmic TDP-43 leads to early functional impairments without neurodegeneration in a Serotonergic Neuron-Specific C. elegans Model. bioRxiv : the preprint server for biology. [PubMed](https://pubmed.ncbi.nlm.nih.gov/40766632/)
[5] Cheemala et al. (2025). Amyotrophic lateral sclerosis and frontotemporal dementia mutation reduces endothelial TDP-43 and causes blood-brain barrier defects. Science advances. [PubMed](https://pubmed.ncbi.nlm.nih.gov/40238886/)
[6] Picher-Martel et al. (2024). TARDBP Mutations in Facial-Onset Sensory and Motor Neuronopathy. Neurology. Genetics. [PubMed](https://pubmed.ncbi.nlm.nih.gov/38841627/)
Similar Research
**Integrative genetic analysis illuminates ALS heritability and identifies risk genes.**
Megat et al. (2023)
*Related research*
[Read on PubMed](https://pubmed.ncbi.nlm.nih.gov/36670122/)
**Biomarker discovery in Alzheimer's and neurodegenerative diseases using Nucleic Acid Linked Immuno-Sandwich Assay.**
Ashton et al. (2025)
*Related research*
[Read on PubMed](https://pubmed.ncbi.nlm.nih.gov/40401628/)
**Frontotemporal dementia. How to deal with its diagnostic complexity?**
Antonioni et al. (2025)
*Related research*
[Read on PubMed](https://pubmed.ncbi.nlm.nih.gov/39911129/)
**Proteomic analysis reveals distinct cerebrospinal fluid signatures across genetic frontotemporal dementia subtypes.**
Sogorb-Esteve et al. (2025)
*Related research*
[Read on PubMed](https://pubmed.ncbi.nlm.nih.gov/39908349/)
**Neuronal dysfunction caused by FUSR521G promotes ALS-associated phenotypes that are attenuated by NF-kappaB inhibition.**
Pelaez et al. (2023)
*Related research*
[Read on PubMed](https://pubmed.ncbi.nlm.nih.gov/37974279/)